Systems Developmental Biology Projects:
Elucidating the mechanism of flux signalling in emergent pattern formation.
 Multiple emergent pattern formation mechnaisms have been suggested to underlie developmental biology phenomena including the Turing-type mechanism (Turing 1952) and the Progressive Oscillatory Reaction Diffusion Mechanism (Cotterell et al., 2015). One major argument often used to rebut the roles of this class of mechanism is there use of diffusion. However, this is really a missunderstanding of what diffusion is representing in these mechanisms. In these mechanism diffusion simply represents 'flux signaling' - i.e. the more of an entity there is over here the less of it there is over there such that mass is conserved. Hence, although we use the word diffusion, this form of signaling could be encoded biologically in multiple different ways. This project aims to elucidate what processes could underlie flux signaling. I have at least two current hypothesis, one involving a particular configuration of cytonemes and another involving a gene product X that undergoes a novel posttranslational moficiation (Left: HEK293 cells transfected with gene X and an IRES to EGFP imaged with STED super resolution microscopy). I aim to study this process in vitro and in the presomitic mesoderm of mouse, chicken and zebrafish embryos.
Multiple emergent pattern formation mechnaisms have been suggested to underlie developmental biology phenomena including the Turing-type mechanism (Turing 1952) and the Progressive Oscillatory Reaction Diffusion Mechanism (Cotterell et al., 2015). One major argument often used to rebut the roles of this class of mechanism is there use of diffusion. However, this is really a missunderstanding of what diffusion is representing in these mechanisms. In these mechanism diffusion simply represents 'flux signaling' - i.e. the more of an entity there is over here the less of it there is over there such that mass is conserved. Hence, although we use the word diffusion, this form of signaling could be encoded biologically in multiple different ways. This project aims to elucidate what processes could underlie flux signaling. I have at least two current hypothesis, one involving a particular configuration of cytonemes and another involving a gene product X that undergoes a novel posttranslational moficiation (Left: HEK293 cells transfected with gene X and an IRES to EGFP imaged with STED super resolution microscopy). I aim to study this process in vitro and in the presomitic mesoderm of mouse, chicken and zebrafish embryos.
Demonstrating the utility of instantaneous nullclines and developing new dynamical systems tools that deal with spatially extended hyperdimensionality
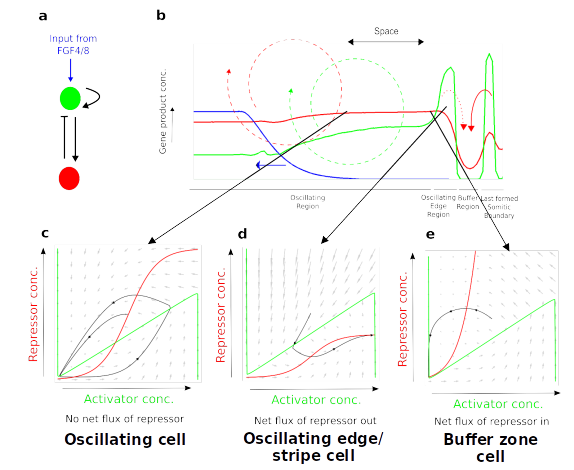
In scenarios where there are multiple components interacting in complex ways intuition typically fails and more advanced tools are requireed to understand a system - this is one of the defining goals of systems biology. The tools of dynamical systems theory including phase spaces and nullclines have proved of great utility for understanding and characterizing the behaviour of biological systems. However, these analytical tools are only applicable to systems with upto about 3 components (at least visually). However, in more complex systems with >3 components or spatially extended systems such as tissues where a gene regulatory network is repeated for each cell with each of those cells signalling to other cells in the environment these analytic tools lose their utility since it is impossible to visualize them in a intuitive way in higher dimensions.
One solution to this problem that i have been developing is called 'instantaneous phase planes/null clines' where one takes advantage of the fact that although the configuration of the phase space/null clines is changing all of the time, there are paricular configurations (a topology of phase space/nullcllines if you like) that accurately represent how the system is behaving as a whole. A good example of this is the instantaneous phase space analysis i have done for the progressive oscillatory reaction diffusion (PORD) mechanism (Cotterell et al., 2015) wheere 3 instantaneous phase spaces/nullclines accurately describe the entire system by describing the 3 basic behavious of the cells (oscillating, frozen peak or frozen trough). The first goal then of this project is to further develop this idea so that it is useful for the systems biology commmunity as a whole.
Secondly, we can take advantage of the fact that although a spatially extended system involves n x c components (where n is the number of components in each cell and c is the number of cells), there is symmetry in the connectivity of the entire system since it involves the same mini complex system repeated many times. The second goal of this project is to develop new dynamical system tools that allow for the visualization of the behaviours of such systems in similar manner to which standard nullcline/phase space anaylsis serves to understand lower complexity systems without having to invoke the 'instantaneous phase space' hack.
|
Disclaimer:
See terms of use tab.
