Omics tool development:
|
| |
|
In order to obtain string evidence for the Emergent theory of cancer model one of the key technologies that needs to be employed is spatial transcriptomics and genomics. Hence, one of the side goals of my laboratory is developing spatial omics techniques that are in general better and more specific for the questions that i need to address. | |
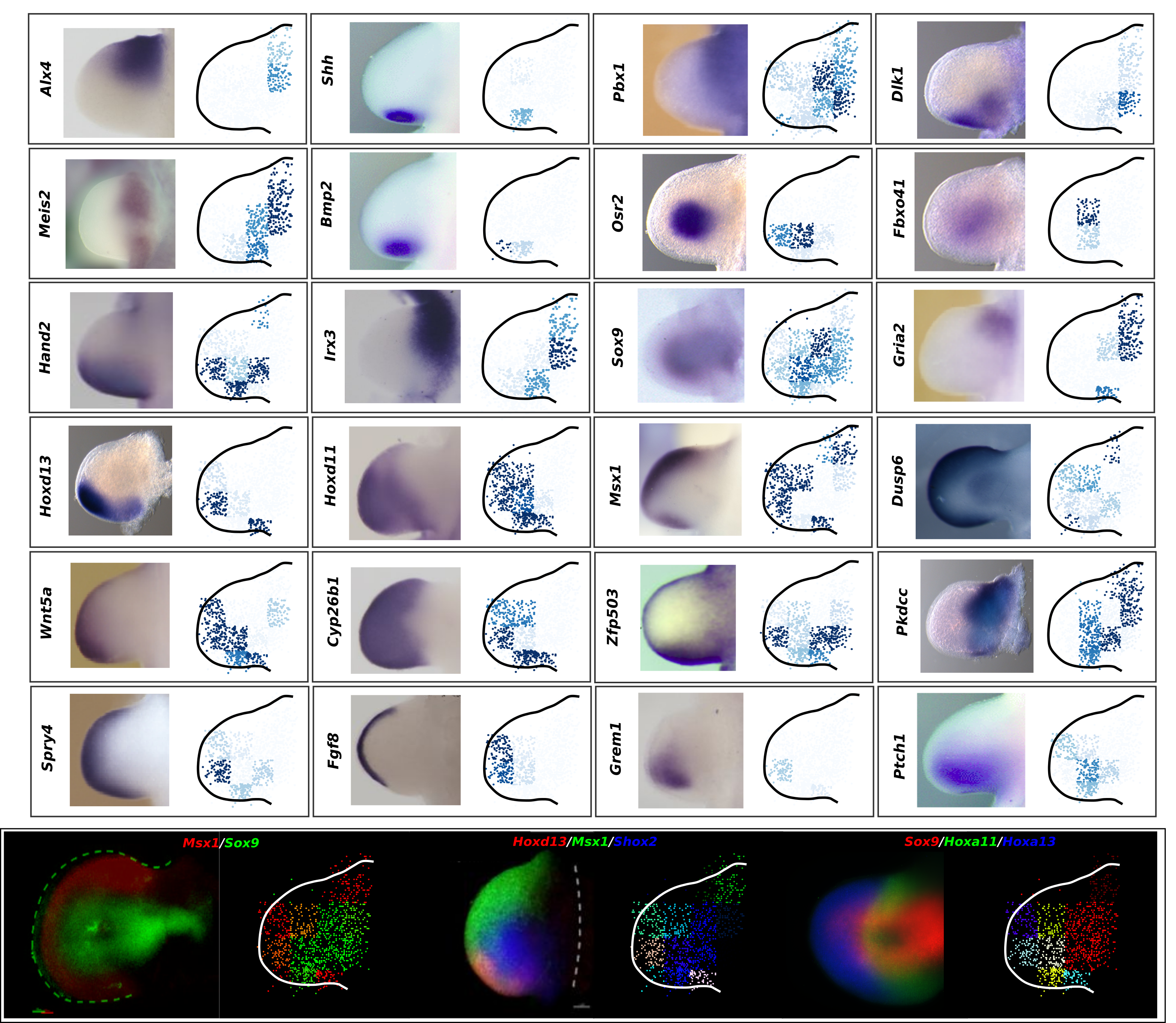
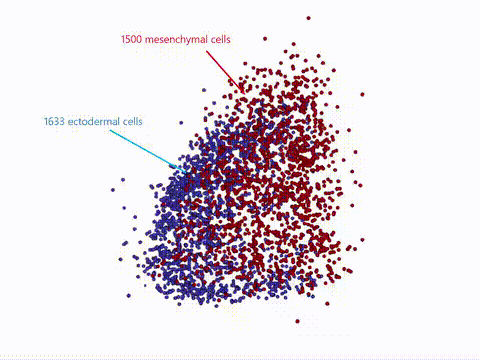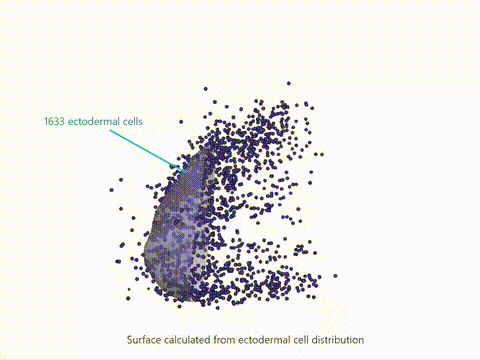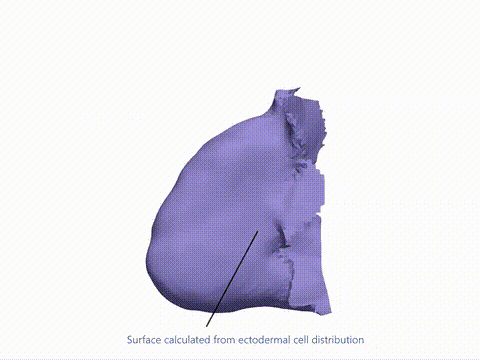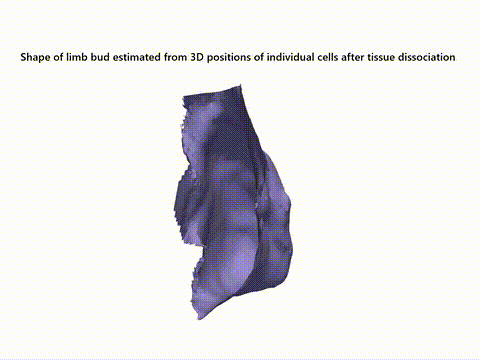

Current state-of-the-art spatial omics approaches suffer from the drawback that they are tissue section-based and thus inherently 2-dimensional. At EMBL Barcelona, over the last several years i have been developing Cell 3D positoning by optical encoding (C3PO), the first technique capable of reconstructing the 3D positions of cells in a tissue, after they have been fully dissociated for single-cell omics analysis (Cotterell et al., 2024). C3PO is not a omics tool per se but is a tool for preserving the positional information of cells once a tissue is dissociated. C3PO imposes a Cartesian coordinate system of positions on the tissue and cells of interest, before dissociation. This is created by multiple orthogonal spatial gradients of active fluorophores, carefully shaped by a 3D bleaching method, such that each position in the tissue is encoded by a unique fluorescent address. Upon dissociation of the tissue the fluorescent addresses of the cells can be read via an appropriate device (such as a FACS machine) to computationally reconstruct the tissue in 3D (right-top), before omics are performed downstream. We have shown proof-of-principle of C3PO for reconstructing the tissue geometry and performing transcriptomics on mouse limb buds (right-bottom: Shows known gene expression patterns as measure by ISH compared to the result obtained by C3PO). I am further developing C3PO in consort with the other members of the C3PO team at EMBL barcelona to improve this technique. | |
|
I am developing alternative technologies to perform 3D spatial transcriptomics and genomics in tissues that are potentially more pragmatic, and have a much higher spatial resolution than C3PO. 1) This technology involves optically writing the spatial DNA barcodes into the tissue such that each cell has a unique spatial barcode. Because there is no optical encoding there is no need for FACS. 2) An alternative Optical encoding technology that like C3PO imposes unique optical barcodes to each of the cells of the tissue but does so in a stochastic manner (Like the brainbow mice but with a lot more information encoded in the hues). I aim to develop these technologies further in this project and apply it to the tumour biology projects i have described above. |
See terms of use tab.
